Identification of bacteria
How do the bacteriologists decide which bacteria is causing the postoperative wound infection?
In about 130 years the bacteriologists used the relatively simple method to smear the pus or the swabs from the wound on to special substrate. The substrate is either fast, gelatin-like substance agar mixed with blood; or liquid, a kind of nutritious soup in a bottle.
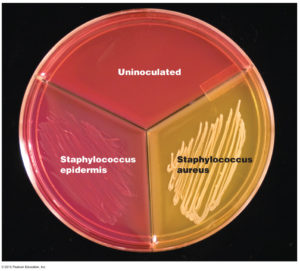
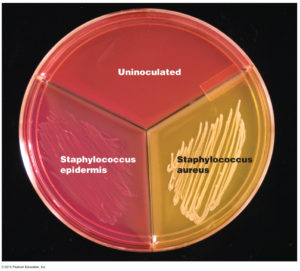
Pus containing Staphylococcus aureus bacteria was smeared on the surface of the blood-agar plate. The blood-agar plate is within a glass dish and is covered by another dish to prevent downfall of other bacteria from the air. The whole culture is grown for at least 2 days in a constant temperature (32-37 degrees Celsius).
After two days the bacteria usually may be identified by growth of characteristic cultures. In this case the cultures of yellow staphylococcus have somehow yellow colour. The bacteriologist can then make microscopic slides from these cultures for direct observation and identification of the bacteria in the microscope (after Gram staining, e.g.).
This method served doctors well for the diagnosis of common infections where the bacteria floated in the pus freely. The scientists call this freely floating bacterial form a planktonic form.
But this method failed for identification of bacteria in torpid, chronic infections of total joints. These infections, that develop very slowly, often grow no bacteria by the usual bacterial cultivation methods.
The reason is that bacteria in these infection adhere firmly to the surface of the total joint and produce a slim cover that protect these bacteria from the action of the antibiotics. The scientists call this bacterial form for sessile bacterial form.
Not only that the slime envelope protects this bacterial form from the direct action of antibiotics, these bacteria successively also are developing extreme resistance against the antibiotics. Moreover, the sessile bacteria do not grow in the laboratory, and thus are impossible to identify by usual bacteriological methods of bacterial identification.
These bacterial forms are, however, possible to identify by two modern methods for direct bacterial identification.
One method is the PCR – polymerase chain reaction method.
This method amplifies a part of bacterial genetic material (RNA) in laboratory. Instead of growing the whole bacteria, which may be impossible, one is “growing” bacteria’s genetic material (RNA) which is quick and easy method done with enzymes. The advantage of this method is its quickness. Simple PCR analysis takes about 13 hours, against 24 to 48 hours (and even more) necessary for conventional bacteriological growth method to identify bacteria.
The disadvantage of this method is its hyper-sensibility. It identifies every bacterium in the studied tissues, even the one that came in the tissue after operation, during transport to the laboratory. Remember that human bacteria are everywhere where are humans.
Because of this fact many bacteriological laboratories abandoned this method for standard identification of bacteria in studied tissues.
There are more specific PCR methods that may exclude this contamination of the studied tissues with “foreign” bacteria; these methods, however, need special laboratory conditions and are as yet very expensive.
Another method is fluorescence microscopy.
The scientist conducts a direct microscopic examination of the bacterial slime layers – films adherent to the removed failed total joints. These slime films are first removed from the surface of the device by ultrasound. Then the bacteria present in these films are stained by specific methods so that they fluorescence. The scientist then studies the slime film by special microscope.
With this method the bacteriologists can easily distinguish the bacteria present in the film from the bacteria that landed (contaminated) in the studied specimen later.
The slime film was stained with specific methods that make the bacteria to fluorescence against a dark background.
The upper picture shows a slime film composed of small round Staphylococcus bacteria. The lower picture shows a slime film composed of the usual skin bacteria living deep in sweat glands with the long name of Propionibacterium acne. This bacterium, our steady companion, is accused of producing torpid total joint infections.
Adapted from McDowell and Patrick 2005
In specialized laboratories the identification of bacteria with the direct microscopic fluorescence method can be made within 3 to 4 hours.
The goal is to make the identification of the bacteria already during the revision operation, within about 30 to 45 minutes.
At present the bacteriologist can study a part of the tissue removed at revision operation for the presence of bacteria by the so called frozen section technique. This simple microscopic examination takes about 15 to 30 minutes, so that the surgeons may have the result during the revision surgery. Unfortunately, this technique misses the bacteria in the tissues – how often? There is a discussion on going on this issue.
From this short overview two things are obvious:
- First: The sensibility of identical bacterial strains to antibiotics differs according to where they are coming from: bacteria that grow on the surface of a total joint have quite different sensitivity to antibiotics than the bacteria causing soft tissue infections. Usually the bacterial strain who is adherent to the surface of a total joint is 1000 or 10 000 times less sensitive to antibiotics than the strain (of the otherwise identical bacteria) that causes soft tissue infections.
- Second: From this fact follows that in chronic infections of total joints the surgeon cannot hope to cure the infection by high doses of antibiotics only. The surgeon must remove the infected total joint device and all soft tissues around it first and then start the antibiotic treatment.
Reference:
- McDowell A, Patrick S.: Evaluation of Nonculture Methods for the Detection of Prosthetic Hip Biofilms. Clin Orthop Rel Research 2005; 437: 74 -82